Record Keeping Helps Bacteria’s Immune System Fight Invaders
Experiments at SLAC that Help Explain Bacterial Adaptive Defenses Could Aid in Disease Prevention
Bacteria have a sophisticated means of defending themselves, and they need it: more viruses infect bacteria than any other biological entity.
Two experiments undertaken at the Department of Energy's SLAC National Accelerator Laboratory provide new insight at the heart of bacterial adaptive defenses in a system called CRISPR, short for Clustered Regularly Interspaced Short Palindromic Repeat.
This portion of bacteria’s immune system works as a record keeper, taking note of attacking viruses’ identities and storing that information by integrating fragments of the virus’ DNA into its own DNA. In this way, CRISPRs maintain genetic records of previously encountered viruses, making it easier for the bacteria’s immune system to send out complexes that destroy viral invaders by identifying and cutting up the recognized DNA sequences.
The studies published last year in Science not only reveal important information about how bacteria repel attacking viruses, but also could potentially improve the prevention of disease in humans. Researchers are currently studying ways of preventing and treating cystic fibrosis, blood disorders and HIV by harnessing the CRISPR system to replace one version of a gene with another or to add a working copy for a mutated gene.
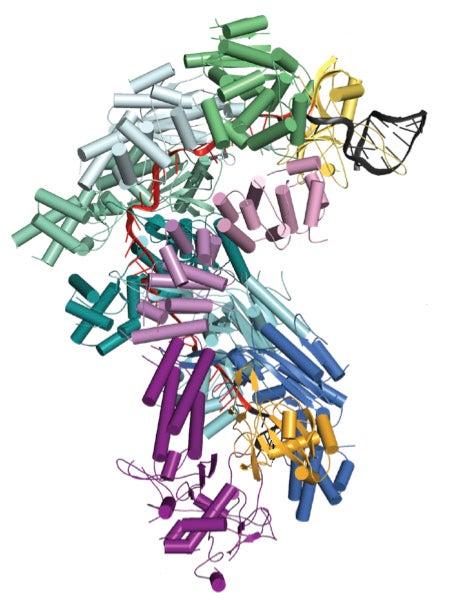
Scientists studied one particular CRISPR-associated complex called Cascade using bright X-rays at SLAC's Stanford Synchrotron Radiation Lightsource (SSRL), a DOE Office of Science User Facility. In the bacteria Escherichia coli, 11 proteins assemble together with an RNA guide that helps Cascade target invading DNA sequences. Once Cascade confirms that the target DNA is from an invader, a molecular signal recruits a nuclease called Cas3 to finish off the invader by chewing it up.
Previous work by Blake Wiedenheft, the Montana State University assistant professor of microbiology and immunology who led one of the studies, and his colleagues revealed Cascade’s seahorse-shaped architecture, but studies undertaken at SSRL now reveal how all the parts of this machine assemble into a functional surveillance machine that patrols the intracellular environment for invading DNA.
“Determining high-resolution structures of large macromolecules remains challenging,” Wiedenheft said. “Several technical aspects of SSRL, including intensity of light, ability to focus the beam, and shutterless X-ray detector made these results possible.”
The studies also revealed that Cascade’s RNA guide does not twist together with the viral DNA to form a helix, as was expected. Instead, they form an under-wound ribbon-like structure.
“A high-resolution structure is essentially a molecular blueprint of a biological machine,” said Wiedenheft. Determining the structure of this complex “is a technical accomplishment that provides the first molecular explanation of how all the parts assemble into a functional surveillance machine.”
Citations: Jackson, R. N. et al. Crystal structure of the CRISPR RNA-guided surveillance complex from Escherichia coli. Science (2014) DOI:10: 10.1126/science.1256328. Mulepati S, et al. Crystal structure of a CRISPR RNA-guided surveillance complex bound to a ssDNA target. Science (2014) 345:1479–1484.
Contact
For questions or comments, contact the SLAC Office of Communications at communications@slac.stanford.edu.
SLAC is a multi-program laboratory exploring frontier questions in photon science, astrophysics, particle physics and accelerator research. Located in Menlo Park, Calif., SLAC is operated by Stanford University for the U.S. Department of Energy's Office of Science.
SLAC National Accelerator Laboratory is supported by the Office of Science of the U.S. Department of Energy. The Office of Science is the single largest supporter of basic research in the physical sciences in the United States, and is working to address some of the most pressing challenges of our time. For more information, please visit science.energy.gov.